Researchers designed a nanoscale “net” that could be used as a drug-delivery system, got help from machine learning to develop a better idea of how RNA works, and took steps toward realizing an immunological holy grail: a universal flu vaccine. Big ideas come in tiny little packages in this week’s coolest scientific findings.
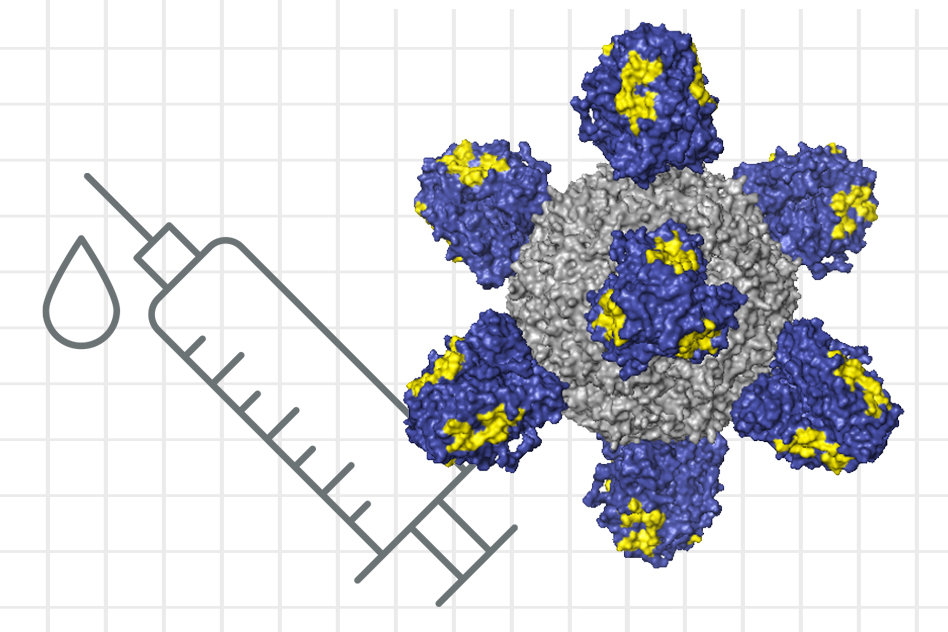
What is it? A collaboration involving researchers from MIT and Harvard has taken steps toward creating a universal flu vaccine.
Why does it matter? A vaccine that confers broad protection from different strains of influenza is, in the words of a 2019 commentary in Nature, a “long-standing dream” among immunologists. Current flu vaccines have to be redesigned every year to account for recent mutations of the virus — and then the public has to be convinced, every year, to get another flu shot. (Which — PSA! — is an especially good idea this year.) MIT professor Arup K. Chakraborty, a senior author of a new study in Cell Systems, said, “The reason we’re excited about this work is that it is a small step toward developing a flu shot that you just take once, or a few times, and the resulting antibody response is likely to protect against seasonal flu strains and pandemic strains as well.”
How does it work? Given the frequent mutation of influenza strains, the challenge is to design a vaccine that teaches the immune system to protect itself against a part of the virus that doesn’t mutate. Made from inactivated viruses, most flu vaccines teach the immune system to target a protein called hemagglutinin, or HA — but the immune system focuses its antibodies on the HA protein’s head, which is the part more prone to mutation than other parts of the protein. So the researchers are seeking, with the help of computer modeling, to direct the immune system toward the stem of the protein, which mutates rarely. Early tests of a potential vaccine in mice have shown promise.

What is it? Researchers at Northwestern University devised a self-assembling nanoscale polymer “net” that could have uses in drug delivery, water purification and medical diagnostics.
Why does it matter? The group’s polymer nanonet captures more than 95% of small molecule drugs, DNA or proteins, “compared to 5%-20% in other delivery systems,” according to the researchers. That ability could be harnessed to, for instance, design vaccines where all molecules have to be delivered simultaneously.
How does it work? The team used a polypropylene sulfone homopolymer, which “forms a wide net through an aqueous solution,” said Evan A. Scott, a professor of biomedical engineering at Northwestern’s McCormick School of Engineering. “Then we induce the net to collapse. It collects anything within the solution, trapping therapeutics inside of nanogel delivery vehicles with very high efficiency.” Postdoctoral fellow Fanfan Du said, “It works like a fishing net, which first spreads out due to electrostatic repulsion and then shrinks upon hydration to trap ‘fish.’” The method is described further in Nature Communications.
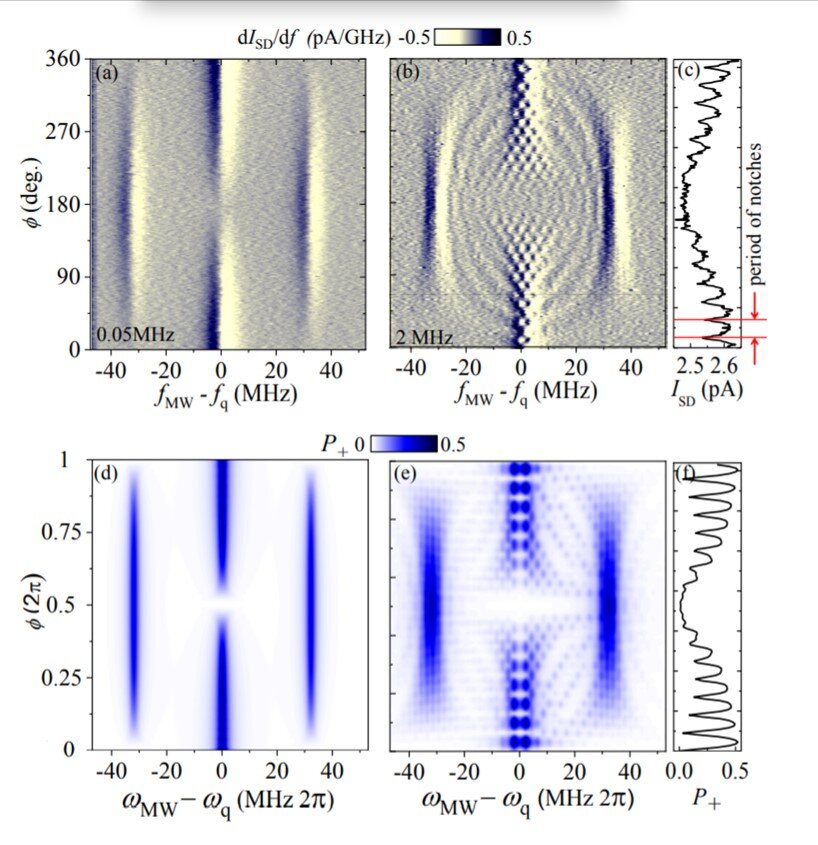
What is it? One of the principles of quantum mechanics — a strange discipline “even physicists don’t understand,” according to a New York Times op-ed — is superposition: the idea that a quantum system can exist in more than one state simultaneously. A collaboration of researchers from Japan, the U.S. and Ukraine recently created a quantum device that illustrates this principle, with the ability to “not only behave analogously to a heat engine and a refrigerator, but also a superposition of both at the same time.”
Why does it matter? Sergey Shevchenko, of Ukraine’s B. Verkin Institute for Low Temperature Physics and Engineering, told Phys.org, “One of the subjects discussed in this field is the possibility of quantum heat engines to overcome the efficiency of classical ones.” While the research is early and the uses of “having a quantum superposition of a tiny engine and a tiny refrigerator” might seem limited, Shevchenko pointed to the example of the laser, which has found myriad surprising applications since its invention: “Possible applications are not clear at this time. This often happens in science.”
How does it work? The basic unit in quantum computing is the qubit, and the challenge Shevchenko and his colleagues faced in exploring thermodynamics and quantum engineering was getting qubits that are “hot, dense and coherent” — which involved a “tunneling field-effect transistor made from densely implanted impurities in silicon.” Read more about the findings here.
What is it? A collaboration between Harvard University’s Wyss Institute and the Massachusetts Institute of Technology used deep learning to better understand how RNA works, which could lead to new RNA-based biological technologies.
Why does it matter? As Wyss explains in a release, DNA and RNA are often compared to “instruction manuals” that govern the systems of our bodies — but unlike, say, computer systems that use binary code, biological systems are “governed by a much messier, more complex set of functions.” This team was interested specifically in the process of RNA folding, said Luis Soenksen, co-first author on one of two new papers in Nature Communications: “These achievements are exciting because they mark the starting point of our ability to ask better questions about the fundamental principles of RNA folding, which we need to know in order to achieve meaningful discoveries and build useful biological technologies.”
How does it work? Soenksen and colleagues focused on RNA “toehold switches,” a type of engineered molecule that, in its “off” state, is folded into a hairpin shape. The way it unfolds allows precise control of gene expression, but researchers have had trouble predicting the behavior of toehold switches. In these studies, the Wyss and MIT teams turned to machine learning to do what it does best: digest massive amounts of information — in this case, a library of nearly 100,000 toehold switches — and learn to detect patterns and predict outcomes. Don Ingber, founding director of the Wyss Institute, said, “The breakthroughs described in these studies demonstrate the power of melding computation with synthetic biology at the bench to develop new and more powerful bioinspired technologies, in addition to leading to new insights into fundamental mechanisms of biological control.”
What is it? An international team led by scientists at the La Jolla Institute for Immunology, the University of Liverpool and the University of Southampton used single-cell RNA sequencing to get the first close look at how the immune system’s CD4+ T cells fight the coronavirus that causes COVID-19.
Why does it matter? As scientists continue to learn how the body reacts to COVID-19 — including the fact that some patients experience the disease much more severely than others — this study drills down on one crucial component of that reaction and establishes a “baseline” for other researchers in their work. “This is the beginning,” said study co-leader Christian Ottensmeier. “We needed to have a reference to look back at for further studies, and this work is novel, timely, detailed, innovative — and open.”
How does it work? The immune system relies on different types of T cells: “helper” cells raise the initial immune alarm about an infection, while other cells make antibodies, and yet other cells act as regulators, making sure the immune system doesn’t go overboard. Using samples from 40 COVID-19 patients — about half who had been hospitalized, with the other half experiencing milder symptoms — researchers analyzed CD4+ T cells as they responded to infection, gaining “a new window into the gene expression patterns that can make each person’s immune response to a virus different,” according to a news release from the La Jolla Institute. The study is described further in the journal Cell.





